Cell Instance Segmentation for Multiple Instance Learning in Digital Pathology¶
Python package for cell instance segmentation and multiple instance learning (MIL) in digital pathology. It provides a complete pipeline from patch extraction to MIL model training and prediction.

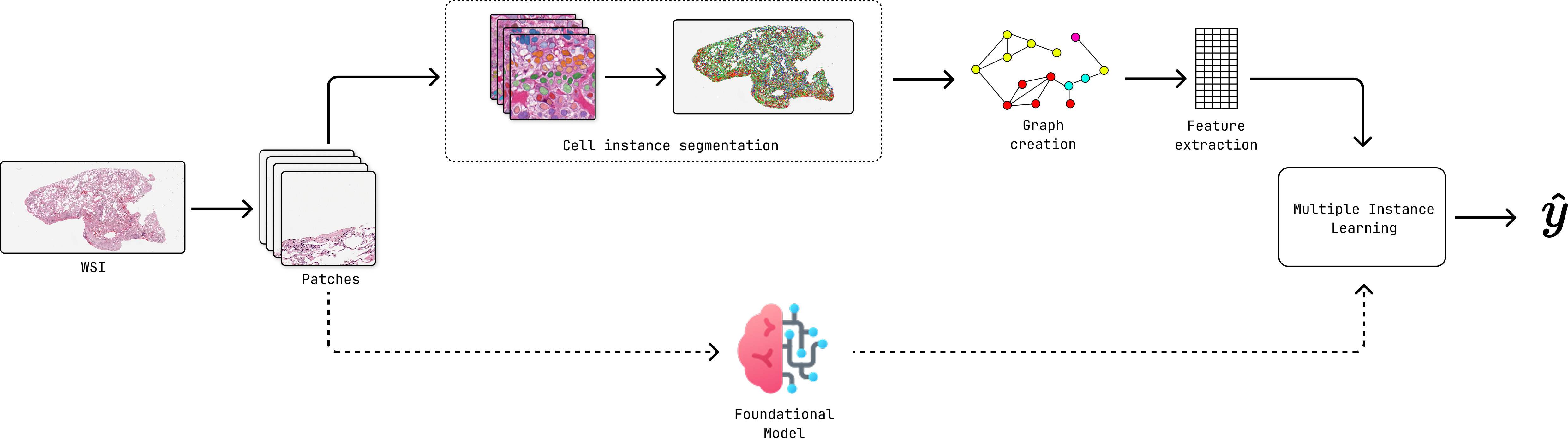
Overview of the cell instance segmentation and multiple instance learning pipeline.¶
Features¶
Patch Extraction: Extract patches from whole slide images (WSI) with configurable parameters
Cell Segmentation: Multiple segmentation models including CellViT, HoverNet, and Cellpose
Graph Creation: Create spatial graphs from segmented cells for improved context understanding
Feature Extraction: Extract morphological and radiomics features from segmented cells
Visualization: Comprehensive feature visualization tools
Quick Start¶
Development Setup¶
For users who want to modify the code or contribute to the project. This is the recommended approach for researchers and developers.
1. Clone the Repository
git clone https://github.com/CamiloSinningUN/CellMIL.git
cd CellMIL
2. Create Conda Environment
Note
This step can be skipped if you already have Python 3.10 and Poetry installed on your machine.
# Create environment
conda env create -f environment.yml
# Activate environment
conda activate cellmil
3. Install Python Dependencies
poetry install
Note
Poetry will install PyTorch with CUDA 11.8 support. If you have a GPU with an older CUDA version, install PyTorch manually first before running poetry install. Visit pytorch.org to get the correct command for your system.
4. Install Additional Dependencies
# Pyradiomics (incompatible with Poetry resolver)
pip install pyradiomics
Note
Optional: Install cucim for GPU-accelerated image loading from their official documentation
Basic Usage¶
Extract patches from WSI:
patch_extraction --output_path ./results --wsi_path ./data/SLIDE_1.svs --patch_size 1024 --patch_overlap 6.25 --target_mag 20.0
Run cell segmentation:
cell_segmentation --model cellvit --gpu 0 --wsi_path ./data/SLIDE_1.svs --patched_slide_path ./results/SLIDE_1
Graph creation:
graph_creation --method delaunay_radius --gpu 0 --patched_slide_path ./results/SLIDE_1 --segmentation_model cellvit
4.1. **Morphological features**:
.. code-block:: bash
feature_extraction --extractor pyradiomics_hed --patched_slide_path ./results/SLIDE_1 --segmentation_model cellvit
4.2. **Topological features**:
.. code-block:: bash
feature_extraction --extractor connectivity --patched_slide_path ./results/SLIDE_1 --segmentation_model cellvit --graph_method delaunay_radius
4.3. **Embedding features**:
.. code-block:: bash
feature_extraction --extractor resnet50 --patched_slide_path ./results/SLIDE_1
Documentation Contents¶
User Guide
API Reference
Development