Model Architectures¶
This section describes available MIL models and when to use them.
Set-Based Models¶
Attention-Based MIL (ABMIL)¶
Ilse, M., Tomczak, J. M., & Welling, M. (2018). Attention-based Deep Multiple Instance Learning.
Classification
from cellmil.models.mil.attentiondeepmil import AttentionDeepMIL, LitAttentionDeepMIL
from torch.optim import AdamW
def lit_model_creator(input_dim: int):
model = AttentionDeepMIL(
embed_dim=input_dim,
size_arg=[256, 128],
n_classes=2,
attention_branches=8,
temperature=1.5,
)
lit_model = LitAttentionDeepMIL(
model=model,
optimizer=AdamW(model.parameters(), lr=1e-4),
)
return lit_model
Survival (SurvAttentionDeepMIL)
from cellmil.models.mil.attentiondeepmil import AttentionDeepMIL, LitSurvAttentionDeepMIL
from cellmil.datamodels.transforms import TimeDiscretizerTransform
from torch.optim import AdamW
label = ("OS_MONTHS", "OS_EVENT")
label_transforms = TimeDiscretizerTransform(n_bins=4)
def lit_model_creator(input_dim: int):
model = AttentionDeepMIL(
embed_dim=input_dim,
size_arg=[256, 128],
n_classes=4, # n_bins
attention_branches=8,
)
lit_model = LitSurvAttentionDeepMIL(
model=model,
optimizer=AdamW(model.parameters(), lr=1e-4),
)
return lit_model
CLAM¶
Data-efficient and weakly supervised computational pathology on whole-slide images. Lu, Ming Y et al., Nature Biomedical Engineering, 2021. DOI: https://doi.org/10.1038/s41551-021-00707-9
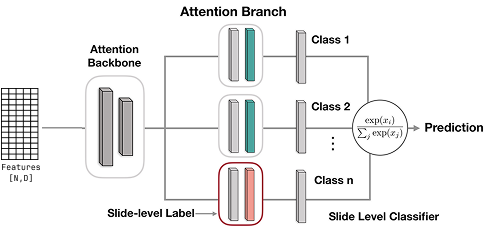
Classification
from cellmil.models.mil.clam import CLAM_SB, LitCLAM
from torch.optim import AdamW
def lit_model_creator(input_dim: int):
model = CLAM_SB(
embed_dim=input_dim,
size_arg="small",
n_classes=2,
k_sample=8
)
lit_model = LitCLAM(
model=model,
optimizer=AdamW(model.parameters(), lr=1e-4),
)
return lit_model
Survival (SurvCLAM)
from cellmil.models.mil.clam import CLAM_SB, LitSurvCLAM
from cellmil.datamodels.transforms import TimeDiscretizerTransform
from torch.optim import AdamW
label = ("OS_MONTHS", "OS_EVENT")
label_transforms = TimeDiscretizerTransform(n_bins=4)
def lit_model_creator(input_dim: int):
model = CLAM_SB(
embed_dim=input_dim,
size_arg="small",
n_classes=4, # n_bins
k_sample=8,
)
lit_model = LitSurvCLAM(
model=model,
optimizer=AdamW(model.parameters(), lr=1e-4),
)
return lit_model
Head4Type¶
Uses separate attention heads for each cell type (neoplastic, inflammatory, connective, etc.). Each cell type gets its own attention mechanism.
Warning
Requirements: This model requires cell type information from the segmentation model. Use CellViT or HoVerNet for segmentation. Cellpose+SAM does not provide cell types and cannot be used with this model.
Note
Don’t use with deep learning features (ResNet50, GigaPath, UNI) as they don’t preserve cell-level type information.
Classification
from cellmil.models.mil.head4type import Head4Type, LitHead4Type
from torch.optim import AdamW
def lit_model_creator(input_dim: int):
model = Head4Type(
embed_dim=input_dim,
size_arg=[256, 128],
n_classes=2,
cell_types=5,
temperature=1.5,
)
lit_model = LitHead4Type(
model=model,
optimizer=AdamW(model.parameters(), lr=1e-4),
)
return lit_model
Survival (SurvHead4Type)
from cellmil.models.mil.head4type import Head4Type, LitSurvHead4Type
from cellmil.datamodels.transforms import TimeDiscretizerTransform
from torch.optim import AdamW
label = ("OS_MONTHS", "OS_EVENT")
label_transforms = TimeDiscretizerTransform(n_bins=4)
def lit_model_creator(input_dim: int):
model = Head4Type(
embed_dim=input_dim,
size_arg=[256, 128],
n_classes=4, # n_bins
cell_types=5,
temperature=1.5,
)
lit_model = LitSurvHead4Type(
model=model,
optimizer=AdamW(model.parameters(), lr=1e-4),
)
return lit_model
CellConv¶
Applies 1D convolutions over the cell sequence before attention pooling. Tries to capture local patterns in spatially-ordered cell features.
It is experimental scenarios only. This model was only used for technical validation while developing CellMIL.
Classification
from cellmil.models.mil.cellconv import CellConv, LitCellConv
from torch.optim import AdamW
def lit_model_creator(input_dim: int):
model = CellConv(
embed_dim=input_dim,
n_classes=2,
convolution_depth=3,
size_arg=[512, 128],
attention_branches=1,
temperature=1.0,
dropout=0.0,
kernel_size=3,
)
lit_model = LitCellConv(
model=model,
optimizer=AdamW(model.parameters(), lr=1e-4),
)
return lit_model
Survival (SurvCellConv)
from cellmil.models.mil.cellconv import CellConv, LitSurvCellConv
from cellmil.datamodels.transforms import TimeDiscretizerTransform
from torch.optim import AdamW
label = ("OS_MONTHS", "OS_EVENT")
label_transforms = TimeDiscretizerTransform(n_bins=4)
def lit_model_creator(input_dim: int):
model = CellConv(
embed_dim=input_dim,
n_classes=4, # n_bins
convolution_depth=3,
size_arg=[512, 128],
attention_branches=1,
)
lit_model = LitSurvCellConv(
model=model,
optimizer=AdamW(model.parameters(), lr=1e-4),
)
return lit_model
Multifocus¶
Uses multi-head attention where each head focuses on different feature dimensions. Aggregates using diagonal elements of the attention-feature product matrix.
It is experimental scenarios only. This model uses a unique diagonal aggregation approach that hasn’t been extensively validated.
from cellmil.models.mil.multifocus import MultiFocus, LitMultiFocus
from torch.optim import AdamW
def lit_model_creator(input_dim: int):
model = MultiFocus(
embed_dim=input_dim,
n_classes=2,
size_arg=[32],
temperature=1.0,
dropout=0.0,
)
lit_model = LitMultiFocus(
model=model,
optimizer=AdamW(model.parameters(), lr=1e-4),
)
return lit_model
Standard MIL¶
Data-efficient and weakly supervised computational pathology on whole-slide images. Lu, Ming Y et al., Nature Biomedical Engineering, 2021. DOI: https://doi.org/10.1038/s41551-021-00707-9
Classification
from cellmil.models.mil.standard import MIL_fc, MIL_fc_mc, LitStandard
from torch.optim import AdamW
def lit_model_creator(input_dim: int):
# Binary classification
model = MIL_fc(
embed_dim=input_dim,
size_arg="small", # or list like [256]
n_classes=2,
top_k=1, # Number of top instances to select
dropout=0.25,
)
# For multi-class (n_classes > 2), use MIL_fc_mc instead:
# model = MIL_fc_mc(embed_dim=input_dim, n_classes=3, top_k=1)
lit_model = LitStandard(
model=model,
optimizer=AdamW(model.parameters(), lr=1e-4),
)
return lit_model
HistoBistro¶
Transformer-based biomarker prediction from colorectal cancer histology: A large-scale multicentric study. Wagner, Sophia J et al., Cancer Cell, Elsevier. DOI: https://doi.org/10.1016/j.ccell.2023.02.002
TransMIL¶
Transmil: Transformer based correlated multiple instance learning for whole slide image classification. Shao, Zhuchen et al., Advances in Neural Information Processing Systems, 2021. DOI: https://proceedings.neurips.cc/paper/2021/hash/10c272d06794d3e5785d5e7c5356e9ff-Abstract.html
Graph-Based Models¶
These models build a graph connecting nearby cells and use graph neural networks to let cells exchange information with their neighbors.
General setup:
from cellmil.datamodels.datasets import GNNMILDataset
# Use GNNMILDataset instead of MILDataset
dataset = GNNMILDataset(
root=config.root,
label=config.label,
folder=config.folder,
data=df,
extractor=config.extractor,
segmentation_model=config.segmentation_model,
graph_creator="delaunay_radius", # Required!
)
Graph Attention Networks (GAT)¶
Veličković, P., Cucurull, G., Casanova, A., Romero, A., Lio, P., & Bengio, Y. (2017). Graph attention networks. arXiv preprint arXiv:1710.10903.
Classification
from cellmil.models.mil.graphmil import GAT, Attention, LitGraphMIL
from cellmil.datamodels.datasets import GNNMILDataset
from cellmil.utils.train.evals import KFoldCrossValidation
from torch.optim import AdamW
# Use GNNMILDataset for graph-based models
dataset = GNNMILDataset(
root=config.root,
label=config.label,
folder=config.folder,
data=df,
extractor=config.extractor,
segmentation_model=config.segmentation_model,
graph_creator="delaunay_radius",
)
def lit_model_creator(input_dim: int):
# GNN backbone
gnn = GAT(
input_dim=input_dim,
hidden_dim=256,
n_layers=4,
dropout=0.25,
heads=3,
)
# Attention pooling head
pooling = Attention(
input_dim=gnn.hidden_dim,
size_arg=[256, 128],
n_classes=2,
attention_branches=8,
temperature=1.5,
)
lit_model = LitGraphMIL(
gnn=gnn,
pooling_classifier=pooling,
optimizer_cls=AdamW,
optimizer_kwargs={"lr": 1e-4},
)
return lit_model
# Run k-fold cross-validation
k_fold = KFoldCrossValidation(k=5)
model_storage = k_fold.evaluate(
name="gat_experiment",
lit_model_creator=lit_model_creator,
dataset=dataset,
output_dir=Path("./results"),
wandb_project="my_project",
)
Survival (SurvGraphMIL)
from cellmil.models.mil.graphmil import GAT, Attention, LitSurvGraphMIL
from cellmil.datamodels.datasets import GNNMILDataset
from cellmil.datamodels.transforms import TimeDiscretizerTransform
from torch.optim import AdamW
label = ("OS_MONTHS", "OS_EVENT")
label_transforms = TimeDiscretizerTransform(n_bins=4)
dataset = GNNMILDataset(
label=label,
graph_creator="delaunay_radius",
# ... other parameters
)
def lit_model_creator(input_dim: int):
gnn = GAT(
input_dim=input_dim,
hidden_dim=256,
n_layers=4,
dropout=0.25,
heads=3,
)
pooling = Attention(
input_dim=gnn.hidden_dim,
size_arg=[256, 128],
n_classes=4, # n_bins
attention_branches=8,
)
lit_model = LitSurvGraphMIL(
gnn=gnn,
pooling_classifier=pooling,
optimizer_cls=AdamW,
optimizer_kwargs={"lr": 1e-4},
)
return lit_model
Other GNN Backbones¶
You can swap GAT with other GNN architectures. All work with both classification (LitGraphMIL) and survival (LitSurvGraphMIL).
GraphSAGE: Hamilton, W., Ying, Z., & Leskovec, J. (2017). Inductive representation learning on large graphs. Advances in neural information processing systems, 30.
from cellmil.models.mil.graphmil import SAGE
gnn = SAGE(
input_dim=input_dim,
hidden_dim=256,
n_layers=3,
dropout=0.25,
)
EGNN (Equivariant GNN): Satorras, V. G., Hoogeboom, E., & Welling, M. (2021, July). E (n) equivariant graph neural networks. In International conference on machine learning (pp. 9323-9332). PMLR.
from cellmil.models.mil.graphmil import EGNN
gnn = EGNN(
input_dim=input_dim,
hidden_dim=256,
n_layers=3,
)
SGFormer: Wu, Q., Zhao, W., Yang, C., Zhang, H., Nie, F., Jiang, H., … & Yan, J. (2023). Sgformer: Simplifying and empowering transformers for large-graph representations. Advances in Neural Information Processing Systems, 36, 64753-64773.
from cellmil.models.mil.graphmil import SGFormer
gnn = SGFormer(
input_dim=input_dim,
hidden_dim=256,
n_layers=3,
)
SmallWorld:
Dynamically creates shortcuts between important cells. Experimental.
from cellmil.models.mil.graphmil import SmallWorld
gnn = SmallWorld(
input_dim=input_dim,
hidden_dim=256,
gamma=0.5,
)
Dynamically creates shortcuts between important cells. Experimental.
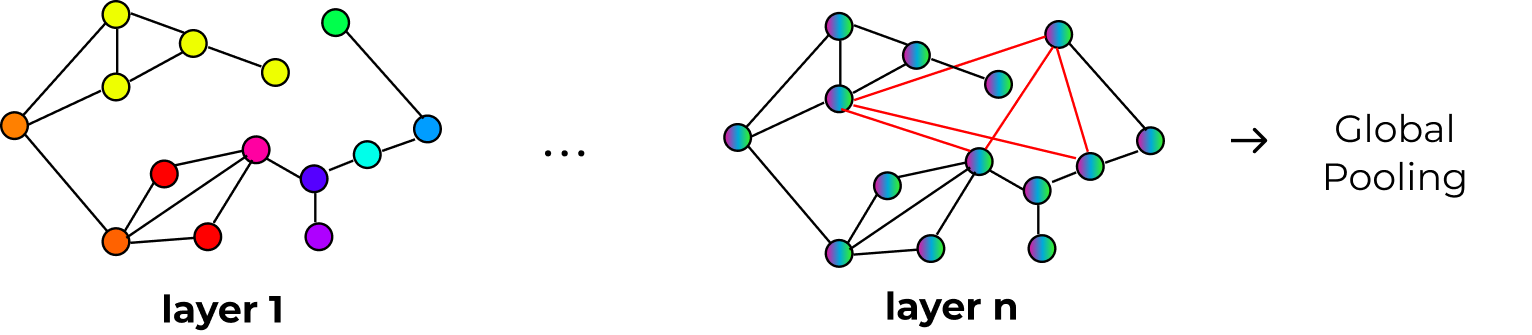
Pooling Methods¶
After the GNN processes the graph, you need to pool node embeddings into a slide-level representation:
Attention Pooling (Recommended):
from cellmil.models.mil.graphmil import Attention
pooling = Attention(
input_dim=gnn.hidden_dim,
size_arg=[256, 128],
n_classes=2,
attention_branches=8,
)
Mean Pooling:
Simple averaging. Use as baseline.
CLAM Pooling:
from cellmil.models.mil.graphmil import CLAM
pooling = CLAM(
input_dim=gnn.hidden_dim,
size_arg=[256, 128],
n_classes=2,
)
Next Steps¶
Training - Train your chosen model
configuration - Configure feature extractors