Cell Segmentation¶
The cell segmentation tool identifies and segments individual cells within extracted patches using deep learning models. This is the second step in the pipeline.
Overview¶
Cell segmentation transforms patch images into cell instance masks and metadata, enabling downstream feature extraction and analysis. The tool supports multiple segmentation models, each optimized for different cell types and imaging conditions.
Available Models¶
CellViT [1]¶
Vision transformer-based model for nuclear instance segmentation and classification.
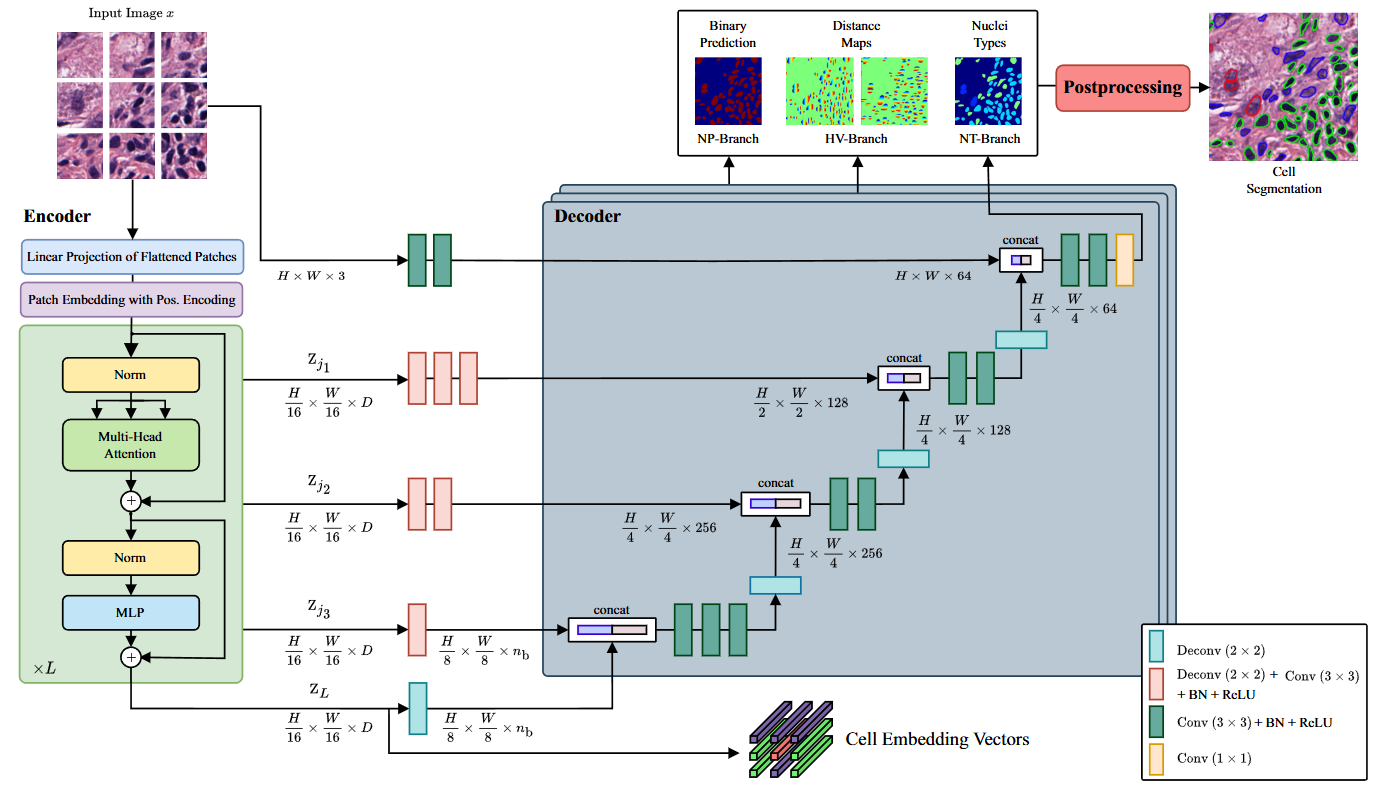
HoVerNet [2]¶
Horizontal and vertical distance regression network for nuclear instance segmentation.
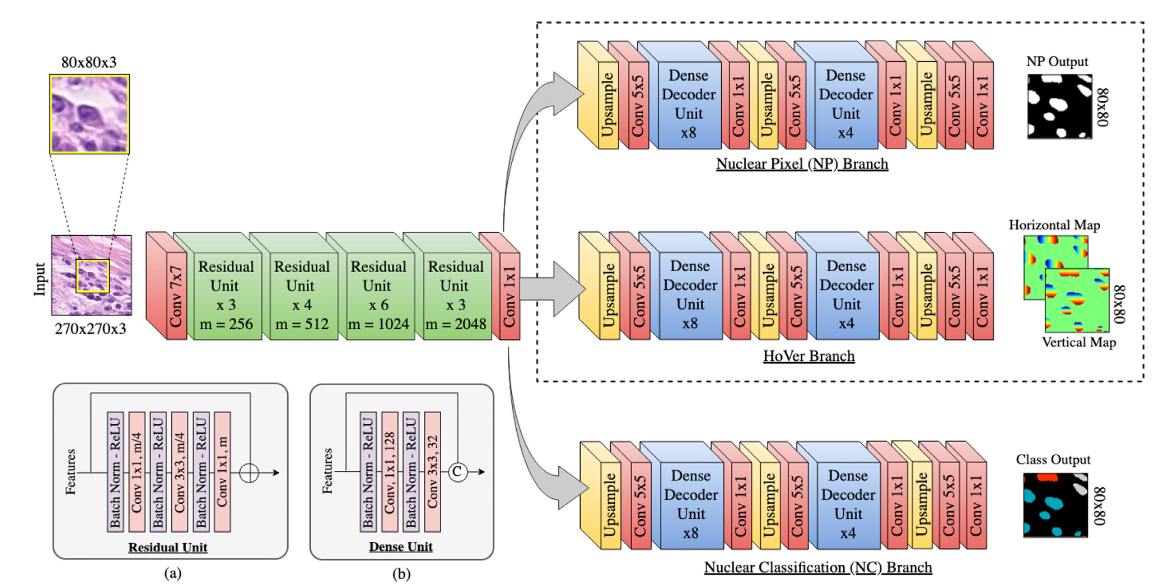
Graham, S., Vu, Q. D., Raza, S. E. A., Azam, A., Tsang, Y. W., Kwak, J. T., & Rajpoot, N. (2019). Hover-net: Simultaneous segmentation and classification of nuclei in multi-tissue histology images. Medical Image Analysis, 101563. https://doi.org/10.1016/j.media.2019.101563
Cellpose + SAM [3]¶
Combination of Cellpose with Segment Anything Model (SAM) for enhanced cell segmentation.
Note
Cell Type Limitation: Cellpose+SAM only provides cell segmentation masks and does not classify cell types. If you need cell type information (required for Head4Type model), use CellViT or HoVerNet instead.
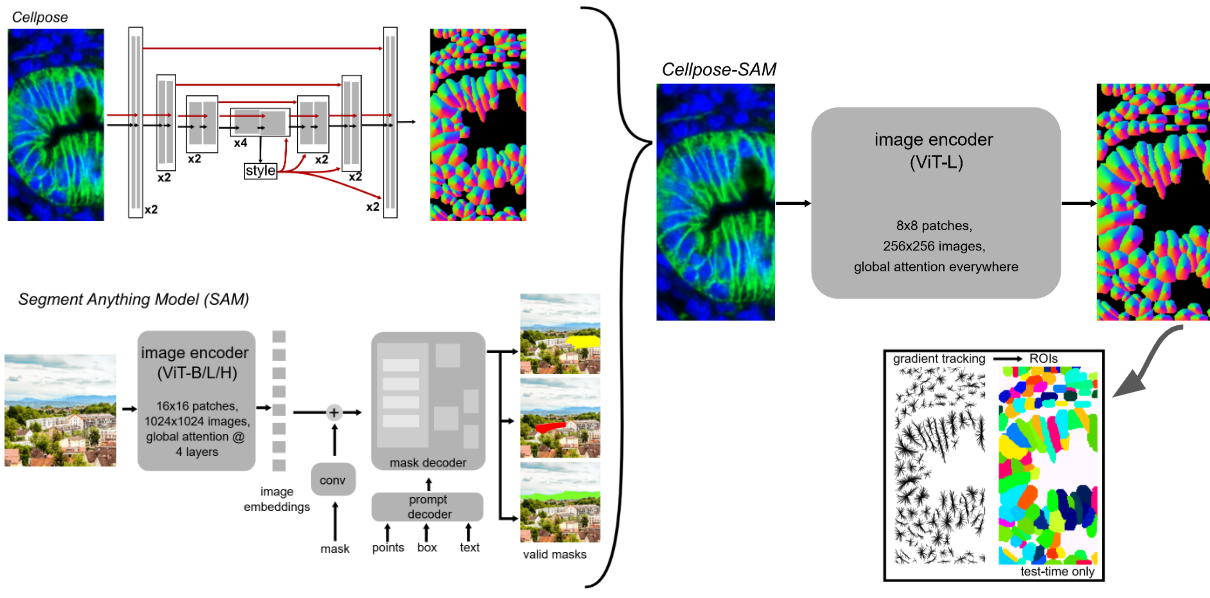
Pachitariu, M., Rariden, M., & Stringer, C. (2025). Cellpose-SAM: superhuman generalization for cellular segmentation. bioRxiv. https://doi.org/10.1101/2025.04.28.651001
CLI Usage¶
Note
⭐ indicates recommended options based on best practices and empirical results.
Basic Command¶
cell_segmentation [OPTIONS]
Required Arguments¶
- --model {cellvit,hovernet,cellpose_sam}¶
Segmentation model to use. Choose based on your specific requirements and cell types.
cellvit: ⭐ Recommended. Provides cell type information.hovernet: Provides cell type information.cellpose_sam: Does not provide cell type information.
- --wsi_path PATH¶
Path to the original whole slide image file. Used for metadata and coordinate mapping.
- --patched_slide_path PATH¶
Path to the directory containing extracted patches (output from patch extraction step).
Optional Arguments¶
- --gpu INTEGER¶
GPU device ID to use for computation.
Default: 0
Complete Example¶
cell_segmentation \
--model cellvit \
--gpu 0 \
--wsi_path ./data/SLIDE_1.svs \
--patched_slide_path ./results/SLIDE_1
This command will:
Load patches from
./results/SLIDE_1/patches/Apply CellViT segmentation model using GPU 0
Generate cell masks and metadata
Save results to
./results/SLIDE_1/cell_detection/cellvit/
Python API Usage¶
You can also perform cell segmentation programmatically:
from cellmil.segmentation import CellSegmenter
from cellmil.interfaces import CellSegmenterConfig
from pathlib import Path
# Create configuration
config = CellSegmenterConfig(
model="cellvit",
gpu=0,
wsi_path=Path("./data/SLIDE_1.svs"),
patched_slide_path=Path("./results/SLIDE_1")
)
# Initialize segmenter
segmenter = CellSegmenter(config)
# Process patches
segmenter.process()
Output Structure¶
Cell segmentation creates the following files:
patched_slide_path/
└── cell_detection/
└── {model_name}/
├── cells.json # Cell instance metadata
├── cells.geojson # Spatial cell data for visualization
├── cell_detection.json # Detection metadata
└── cell_detection.geojson # Spatial detection data
File Descriptions¶
- cells.json
Contains cell instance information including:
Cell boundaries and centroids
Patch coordinates and mappings
Classification scores (if applicable)
- cells.geojson
Spatial representation of cells in GeoJSON format for visualization in tools like QuPath.
- cell_detection.json
Detection-level metadata including:
Detection confidence scores
Centroids and bounding boxes
Quality Assessment¶
Visual Inspection¶
After segmentation, inspect results by:
Loading GeoJSON files in QuPath or similar viewer
Checking cell detection accuracy on representative patches
Verifying segmentation quality in different tissue regions
Integration with Pipeline¶
Cell segmentation output is used by:
Feature Extraction: Extract morphological features from segmented cells
Graph Creation: Create spatial graphs from segmented cells
The standardized output format ensures compatibility with downstream tools.
See Also¶
Patch Extraction - Previous step in the pipeline
Graph Creation - Next step in the pipeline
Feature Extraction - Next step in the pipeline
cellmil.segmentation - API documentation
Quick Start - Complete pipeline overview