Tissue Detection¶
Tissue detection identifies regions containing tissue vs. background in whole slide images. This is a critical preprocessing step in digital pathology pipelines.
Important
The built-in OtsuTissueDetector is a reference
implementation meant to demonstrate the TissueDetector
interface. For production use, we strongly recommend implementing your own detector tailored
to your specific tissue types, staining protocols, and scanning artefacts.
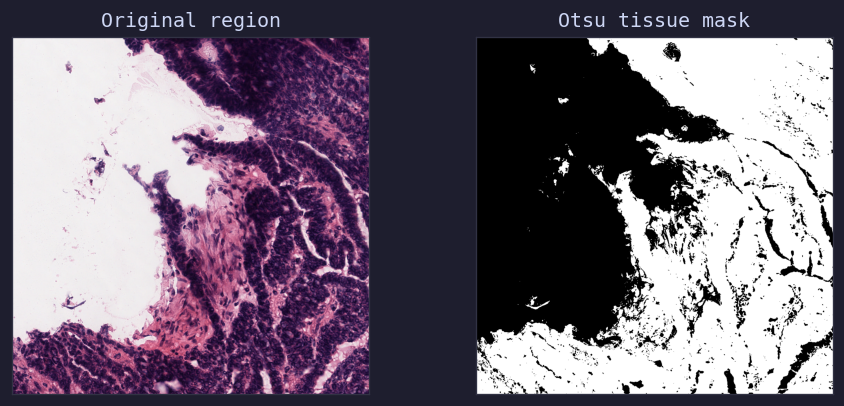
Otsu thresholding on heart tissue — original region, binary tissue mask, and overlay¶
OtsuTissueDetector¶
from glasscut import OtsuTissueDetector
from PIL import Image
detector = OtsuTissueDetector()
image = Image.open("region.png")
mask = detector.detect(image) # np.ndarray, 0=background, 1=tissue
The detector converts the image to grayscale, applies Otsu’s thresholding via
skimage.filters.threshold_otsu, and returns a binary mask.
Custom Detectors¶
Implement a custom tissue detector by subclassing TissueDetector:
from glasscut.tissue_detectors import TissueDetector
import numpy as np
from PIL import Image
class MyTissueDetector(TissueDetector):
def detect(self, image: Image.Image) -> np.ndarray:
# Your custom detection logic here
# Return binary mask: 0 = background, 1 = tissue
pass
Using with GridTiler¶
Pass your detector instance to the tiler:
tiler = GridTiler(
tissue_detector=MyTissueDetector(),
min_tissue_ratio=0.2,
)