Stain Normalization¶
Stain normalization reduces colour variation caused by differences in staining protocols, scanners, and laboratory conditions across histopathology slides.
Important
The built-in MacenkoStainNormalizer and
ReinhardtStainNormalizer are reference
implementations provided to demonstrate the StainNormalizer
interface. Both emit a UserWarning indicating their experimental status.
For production use, we strongly recommend implementing your own normalizer adapted to your specific stain types, tissue morphology, and computational constraints.
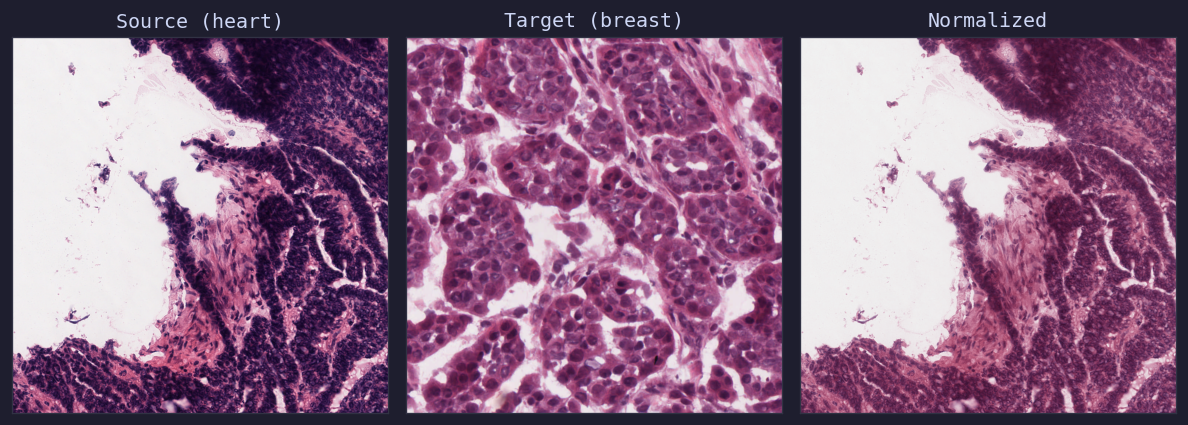
Macenko normalization: heart tissue (source) matched to breast tissue (target)¶
Macenko Normalizer¶
The Macenko method uses principal component analysis (PCA) on optical density values to estimate stain vectors, then matches the stain distribution to a target image.
from glasscut import MacenkoStainNormalizer
from PIL import Image
normalizer = MacenkoStainNormalizer()
target = Image.open("target.png")
normalizer.fit(target)
source = Image.open("source.png")
normalized = normalizer.transform(source)
normalized.save("normalized.png")
Warning
The Macenko normalizer is marked as experimental. Results may vary across different tissue types and staining qualities.
Reinhardt Normalizer¶
The Reinhardt method matches the mean and standard deviation of each LAB colour channel on tissue regions, providing a fast approximate normalisation.
from glasscut import ReinhardtStainNormalizer
from PIL import Image
normalizer = ReinhardtStainNormalizer()
target = Image.open("target.png")
normalizer.fit(target)
source = Image.open("source.png")
normalized = normalizer.transform(source)
normalized.save("normalized.png")
Warning
The Reinhardt normalizer is marked as experimental. Results may vary across different tissue types and staining qualities.
Custom Normalizers (Recommended)¶
Implement a custom normalizer by subclassing StainNormalizer:
from glasscut.stain_normalizers import StainNormalizer
from PIL import Image
class MyNormalizer(StainNormalizer):
def fit(self, target_image: Image.Image, **kwargs):
pass
def transform(self, image: Image.Image, **kwargs) -> Image.Image:
pass
Using with GridTiler¶
Stain normalizers can be used as tile transforms in the tiling pipeline:
normalizer = MacenkoStainNormalizer()
normalizer.fit(target_image)
tiler = GridTiler(
tile_size=(512, 512),
transforms=[normalizer.transform],
)